Biology and Biophysics of Nanoscale Bioparticles
Nanoscale bioparticles such as exosomes, lipoproteins and lipid nanoparticles are crucial for health. Surface, content and biophysical properties of these particles in the body can be used as early biomarkers for diseases. Despite their central roles in physiology, our understanding of the precise biology and structural/functional heterogeneity of these particles is very limited. Available methodologies, that are bulk and low throughput, do not allow us to study different types of nanoscale bioparticles present in complex milieu such as blood, each presumably fulfilling a different function.
We develop and apply novel tools to reveal the heterogeneity of thousands of nanoscale bioparticles in terms of amount, properties and content in a fast and a high throughput manner.
Key publications: Sych et al, Nature Biotech, 2023; Wiklander et al, Nature Biomedical Engineering, 2024.
Nanoscale bioparticles such as exosomes, lipoproteins and lipid nanoparticles are crucial for health. Surface, content and biophysical properties of these particles in the body can be used as early biomarkers for diseases. Despite their central roles in physiology, our understanding of the precise biology and structural/functional heterogeneity of these particles is very limited. Available methodologies, that are bulk and low throughput, do not allow us to study different types of nanoscale bioparticles present in complex milieu such as blood, each presumably fulfilling a different function.
We develop and apply novel tools to reveal the heterogeneity of thousands of nanoscale bioparticles in terms of amount, properties and content in a fast and a high throughput manner.
Key publications: Sych et al, Nature Biotech, 2023; Wiklander et al, Nature Biomedical Engineering, 2024.
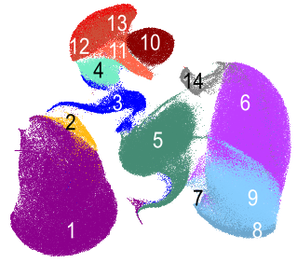
Physics of cells in health and diseases
Critical to cellular function are the collective biophysical and physicochemical properties, such as stiffness, fluidity, and viscosity. There is a compelling need to comprehensively map these properties in health and disease; and exploit them as biomarkers for various cell states. However, conventional methodologies for measuring these properties have traditionally suffered from low throughput, significantly impeding the translation of such concepts into practical applications within the realms of biomedicine.
We fill these gaps by developing and applying novel methods designed for the high throughput and single-cell measurement of biophysical properties across multiple cell types and states simultaneously.
Key publications: Andronico et al, Biorxiv 2024
Critical to cellular function are the collective biophysical and physicochemical properties, such as stiffness, fluidity, and viscosity. There is a compelling need to comprehensively map these properties in health and disease; and exploit them as biomarkers for various cell states. However, conventional methodologies for measuring these properties have traditionally suffered from low throughput, significantly impeding the translation of such concepts into practical applications within the realms of biomedicine.
We fill these gaps by developing and applying novel methods designed for the high throughput and single-cell measurement of biophysical properties across multiple cell types and states simultaneously.
Key publications: Andronico et al, Biorxiv 2024
Biophysics of Immune Cell Signalling
The immune system protects our bodies from extrinsic as well as intrinsic sources of harm. Malfunction of immune cells leads to serious disorders such as cancers, infections, and autoimmune diseases. Key to the success of the immune system are the complex yet perfectly coordinated signalling mechanisms that immune cells use to distinguish friend from foe, guiding their attacks on external pathogens and malfunctioning endogenous components (e.g. cancer cells). Thus, understanding the mechanisms of immune signalling is essential for development of efficient therapeutic approaches against cancer, infection, and auto-immune diseases.
We aim to understand the physical principles underlying immune signalling.
Our Collaborators: Mike Dustin (Oxford), Petter Brodin (Karolinska), Bjorn Onfelt (Karolinska), Evren Alici (Karolinska)
Key publications: Jenkins et al, JCS, 2018; Felce&Sezgin et al, Science Sig, 2018
The immune system protects our bodies from extrinsic as well as intrinsic sources of harm. Malfunction of immune cells leads to serious disorders such as cancers, infections, and autoimmune diseases. Key to the success of the immune system are the complex yet perfectly coordinated signalling mechanisms that immune cells use to distinguish friend from foe, guiding their attacks on external pathogens and malfunctioning endogenous components (e.g. cancer cells). Thus, understanding the mechanisms of immune signalling is essential for development of efficient therapeutic approaches against cancer, infection, and auto-immune diseases.
We aim to understand the physical principles underlying immune signalling.
Our Collaborators: Mike Dustin (Oxford), Petter Brodin (Karolinska), Bjorn Onfelt (Karolinska), Evren Alici (Karolinska)
Key publications: Jenkins et al, JCS, 2018; Felce&Sezgin et al, Science Sig, 2018
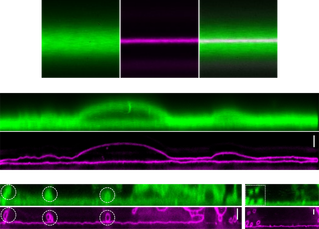

Development of Super-Resolution Techniques
Signalling processes at the plasma membrane occur at very fast temporal and small spatial scales. Therefore, studies aimed at a thorough molecular understanding of cell signalling require advanced super-resolution techniques with nanometre and microseconds spatiotemporal resolution that can capture the molecular interactions.
We constanty develop, modify and apply advanced imaging and spectroscopy tools to reveal the nanoscale mysteries of cells.
Our Collaborators: Ilaria Testa (KTH), Hjalmar Brismar (KTH), Hans Blom (KTH)
Key publications: Barbotin et al, Biophy Jour, 2020; Sezgin et al, Biophy Jour, 2017; Schneider et al, Nano Letters, 2018; Sezgin et al, Nature Protocols, 2019
Signalling processes at the plasma membrane occur at very fast temporal and small spatial scales. Therefore, studies aimed at a thorough molecular understanding of cell signalling require advanced super-resolution techniques with nanometre and microseconds spatiotemporal resolution that can capture the molecular interactions.
We constanty develop, modify and apply advanced imaging and spectroscopy tools to reveal the nanoscale mysteries of cells.
Our Collaborators: Ilaria Testa (KTH), Hjalmar Brismar (KTH), Hans Blom (KTH)
Key publications: Barbotin et al, Biophy Jour, 2020; Sezgin et al, Biophy Jour, 2017; Schneider et al, Nano Letters, 2018; Sezgin et al, Nature Protocols, 2019
Development of Synthetic Biology Tools
Thorough investigation of cell signalling has so far been hampered by the complex structure of the cell membrane. To clarify the biophysical and physicochemical principles underlying cell signalling at the molecular level and discern the essential drivers of the processes, well-defined synthetic biology tools with controllable molecular specificity and complexity are required.
We develop bottom-up and top-down synthetic biology tools to pinpoint the physicochemical principles underlying biological processes. Our tools span from lipid-only systems, to model vesicles with specific lipids and proteins, to plasma membrane vesicles with modular reconstitution of proteins, cytoskeleton and energetic processes, to finally live primary cells.
Key publications: Sezgin et al, Nature Protocols, 2012; Cespedes et al, FEBS Journal, 2019
Thorough investigation of cell signalling has so far been hampered by the complex structure of the cell membrane. To clarify the biophysical and physicochemical principles underlying cell signalling at the molecular level and discern the essential drivers of the processes, well-defined synthetic biology tools with controllable molecular specificity and complexity are required.
We develop bottom-up and top-down synthetic biology tools to pinpoint the physicochemical principles underlying biological processes. Our tools span from lipid-only systems, to model vesicles with specific lipids and proteins, to plasma membrane vesicles with modular reconstitution of proteins, cytoskeleton and energetic processes, to finally live primary cells.
Key publications: Sezgin et al, Nature Protocols, 2012; Cespedes et al, FEBS Journal, 2019
Cell Membrane Structure and Dynamics
The cellular plasma membrane is a heterogeneous structure composed of various types of lipids and proteins, and this heterogeneity plays crucial roles in cellular signalling. The underlying physicochemical principles have been extensively studied for many years, however our understanding of cell memrbane structure is fairly limited.
By applying above mentioned technologies as well as biochemical methods and lipidomics, we aim to elucidate how the cell membrane is organised at nanoscale. We are particularly interested in lipid-lipid and lipid-protein interactions at the plasma membrane and their role in cellular physiology.
Our Collaborators: Ilya Levental (Texas), Ed Lyman (Delaware)
Key publications: Pinkwart et al, JBC, 2019; Sezgin et al, Nature Reviews, 2017
The cellular plasma membrane is a heterogeneous structure composed of various types of lipids and proteins, and this heterogeneity plays crucial roles in cellular signalling. The underlying physicochemical principles have been extensively studied for many years, however our understanding of cell memrbane structure is fairly limited.
By applying above mentioned technologies as well as biochemical methods and lipidomics, we aim to elucidate how the cell membrane is organised at nanoscale. We are particularly interested in lipid-lipid and lipid-protein interactions at the plasma membrane and their role in cellular physiology.
Our Collaborators: Ilya Levental (Texas), Ed Lyman (Delaware)
Key publications: Pinkwart et al, JBC, 2019; Sezgin et al, Nature Reviews, 2017

